


1/12

品牌: CST
 下载产品说明书
下载产品说明书 用小程序,查商品更便捷
用小程序,查商品更便捷



 收藏
收藏
 对比
对比 咨询
咨询
产品介绍
产品信息
简单描述
CUT&RUN protein-DNA interaction assay overcomes many of the challenges faced by WGS mapping techniques. Fast, low sample & cost effective. Click.

组合货号
72917P&91931P

应用
目标/特异性
Specificity/Sensitivity
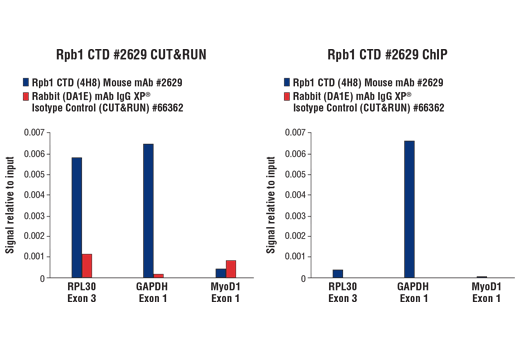
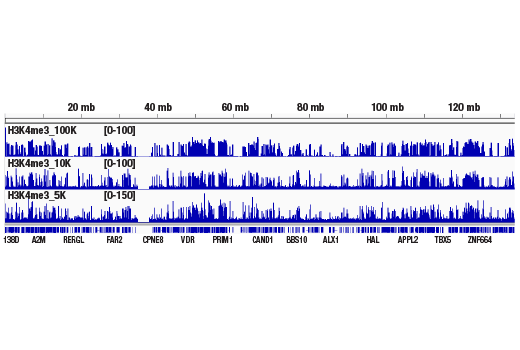
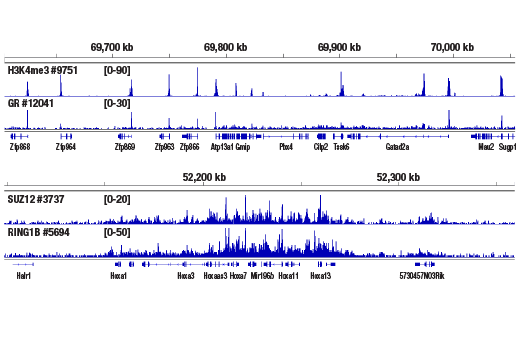
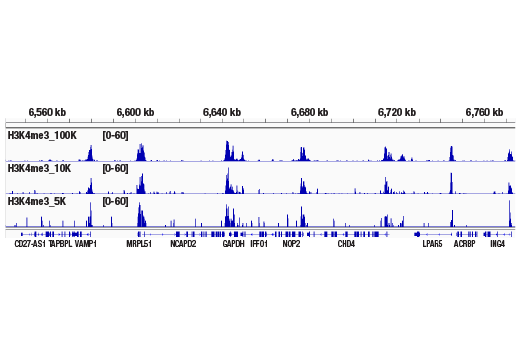
The CUT&RUN Assay Kit can be utilized with any CUT&RUN-validated antibody to detect endogenous levels of protein-DNA interactions and histone modifications in mammalian cells (see Figures 1–6). The kit is compatible with multiple species of antibodies, including rabbit and mouse. The positive control Tri-Methyl-Histone H3 (Lys4) (C42D8) Rabbit mAb #9751 detects multiple species of tri-methyl histone H3 Lys4 protein, including human, mouse, rat, and monkey. Primer sets are included for the human (#7014) and mouse (#7015) positive control RPL30 gene locus; however, the use of other species with the kit requires the design of additional control primer sets.

背景
背景
Like the chromatin immunoprecipitation (ChIP) assay, Cleavage Under Targets & Release Using Nuclease (CUT&RUN) is a powerful and versatile technique used for probing protein-DNA interactions within the natural chromatin context of the cell (1-4). This assay can be used to identify multiple proteins associated with a specific region of the genome, or the opposite, to identify the many regions of the genome associated with a particular protein. In addition, the CUT&RUN assay can be used to define the spatial and temporal relationship of a particular protein-DNA interaction. For example, the CUT&RUN assay can be used to determine the specific order of recruitment of various protein factors to a gene promoter or to “measure” the relative amount of a particular histone modification across an entire gene locus during gene activation. In addition to histone proteins, the CUT&RUN assay can also be used to analyze binding of transcription factors and cofactors, DNA replication factors, and DNA repair proteins (Figures 1-6).CUT&RUN provides a rapid, robust, and true low cell number assay for detection of protein-DNA interactions in the cell. Unlike the ChIP assay, CUT&RUN is free from formaldehyde cross-linking, chromatin fragmentation, and immunoprecipitation, making it a much faster and more efficient method for enriching protein-DNA interactions and identifying target genes. CUT&RUN can be performed in less than one day, from live cells to purified DNA, and has been shown to work with as few as 500-1000 cells per assay (1,2). Instead of fragmenting all of the cellular chromatin as done in ChIP, CUT&RUN utilizes an antibody-targeted digestion of chromatin, resulting in much lower background signal than seen in the ChIP assay. As a result, CUT&RUN requires only 1/10th of the sequencing depth that is required for ChIP-seq assays (1,2). Finally, the inclusion of simple spike-in control DNA allows for accurate quantification and normalization of target-protein binding that is not possible with the ChIP method. This provides for effective normalization of signal between samples and between experiments.
1.Skene, P.J. and Henikoff, S. (2017) Elife 6, pii: e21856. doi: 10.7554/eLife.21856.
2.Skene, P.J. et al. (2018) Nat Protoc 13, 1006-19.
3.Meers, M.P. et al. (2019) Elife 8, pii: e46314. doi: 10.7554/eLife.46314.
4.Meers, M.P. et al. (2019) Mol Cell 75, 562-575.e5.

制备和贮存
保存方式
All components in this kit are stable for at least 12 months when stored at the recommended temperature.
声明 :本官网所有报价均为常温或者蓝冰运输价格,如有产品需要干冰运输,需另外加收干冰运输费。












 危险品化学品经营许可证(不带存储) 许可证编号:沪(杨)应急管危经许[2022]202944(QY)
危险品化学品经营许可证(不带存储) 许可证编号:沪(杨)应急管危经许[2022]202944(QY)  营业执照(三证合一)
营业执照(三证合一)